
Ronald Gieschke, Robert Carr
November 7, 2022
CPT: Pharmacometrics & Systems Pharmacology
Selected publications from ISB are listed below.

Ronald Gieschke, Robert Carr
Manuela Cassotta, Hugo Geerts, Lise Harbom, Tiago F. Outeiro, Iosif Pediaditakis, Orly Reiner, Stefan Schildknecht, Jens C. Schwamborn, Jarrod Bailey, Kathrin Herrmann, Helena T. Hogberg
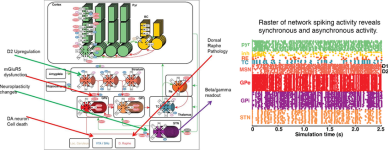
Rachel Rose, Emma Mitchell, Piet Van Der Graaf, Daisuke Takaichi, Jun Hosogi, Hugo Geerts
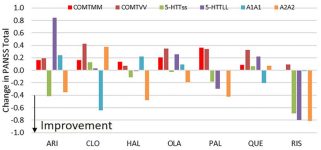
Athan Spiros and Hugo Geerts

Peter Bloomingdale, Tatiana Karelina, Murat Cirit, Sarah F. Muldoon, Justin Baker, William J. McCarty, Hugo Geerts, Sreeraj Macha
Hugo Geerts

Petri Takkala and Steven A. Prescott
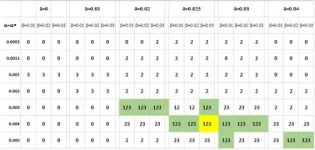
Hugo Geerts, Athan Spiros, Patrick Roberts
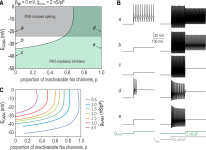
Petri Takkala, Yi Zhu, Steven A. Prescott
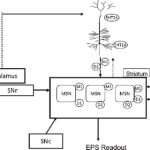
Athan Spiros, Patrick Roberts, Hugo Geerts